Saturday 22 February 2025
Researchers have developed a novel approach to analyzing microbiome data, which could revolutionize our understanding of the intricate relationships between different microorganisms in our bodies.
The human microbiome is composed of trillions of microorganisms that live inside and on our bodies. These microbes play a crucial role in our health, influencing everything from digestion to immunity. However, understanding their complex interactions has been a significant challenge due to the vast amounts of data generated by high-throughput sequencing technologies.
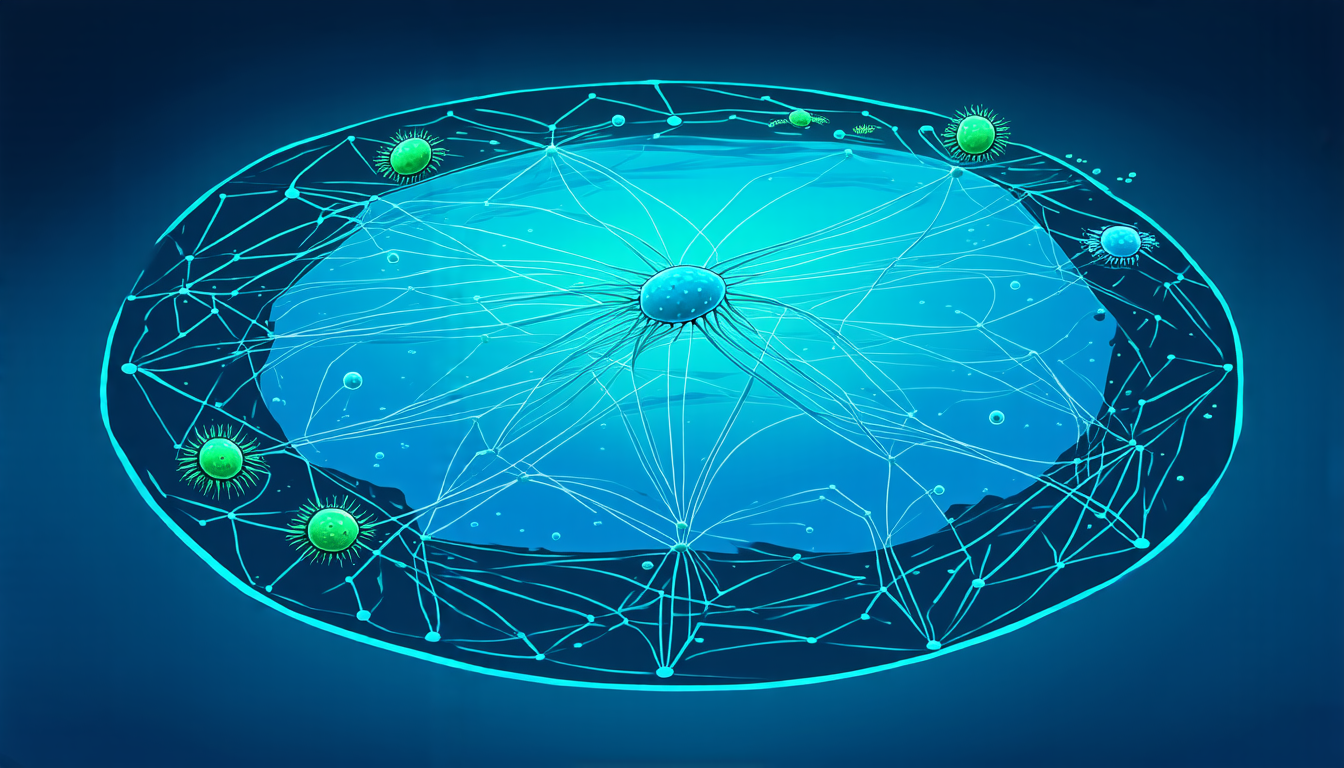
To tackle this issue, scientists have developed a new method for inferring co-occurrence networks from microbiome data. This approach involves processing raw genetic reads and constructing networks that describe the ecosystems of different body sites under various conditions.
The researchers tested their methodology on a dataset of chickens vaccinated against and challenged by the protozoan parasite Eimeria tenella. They found that their method was able to extract biologically meaningful information from the data, including the evolution of node distributions over time.
One key advantage of this approach is its ability to handle small sample sizes, which are common in many biological studies. The method uses a combination of statistical filters and network analysis techniques to identify significant patterns and relationships in the data.
The implications of this research are far-reaching. For example, it could be used to develop personalized treatment strategies for diseases that involve alterations to the gut microbiome, such as irritable bowel syndrome or inflammatory bowel disease.
Furthermore, this approach could also be applied to other types of omics data, including transcriptomics and proteomics. By integrating these different datasets, scientists may gain a more comprehensive understanding of biological systems and develop new treatments for complex diseases.
Overall, this research demonstrates the power of graph-based machine learning methods in microbiome analysis and highlights their potential to transform our understanding of microbial ecosystems.
Cite this article: “Deciphering Microbiome Complexities with Graph-Based Machine Learning”, The Science Archive, 2025.
Microbiome, Machine Learning, Graph-Based, Co-Occurrence Networks, High-Throughput Sequencing, Data Analysis, Statistical Filters, Node Distributions, Personalized Treatment, Omics Data